The Basic Energy Sciences (BES) program supports basic scientific research to lay the foundations for new energy technologies and to advance DOE missions in energy, environment, and national security. BES research emphasizes discovery, design, and understanding of new materials and new chemical, biochemical, and geological processes. The ultimate goal is to better understand the physical world and harness nature to benefit people and society.
Major technological innovations don’t just happen. They typically have their roots in basic research breakthroughs over a period of decades. The BES program supports basic research behind a broad range of energy technologies, spanning energy generation, conversion, transmission, storage, and use. Many major innovations can be traced back to basic research supported by BES over the past 40 years. These include, for example, LED lighting; efficient solar cells; better batteries; stronger, lighter materials for transportation, nuclear power plants, and national defense; and improved production processes for high-value chemicals.
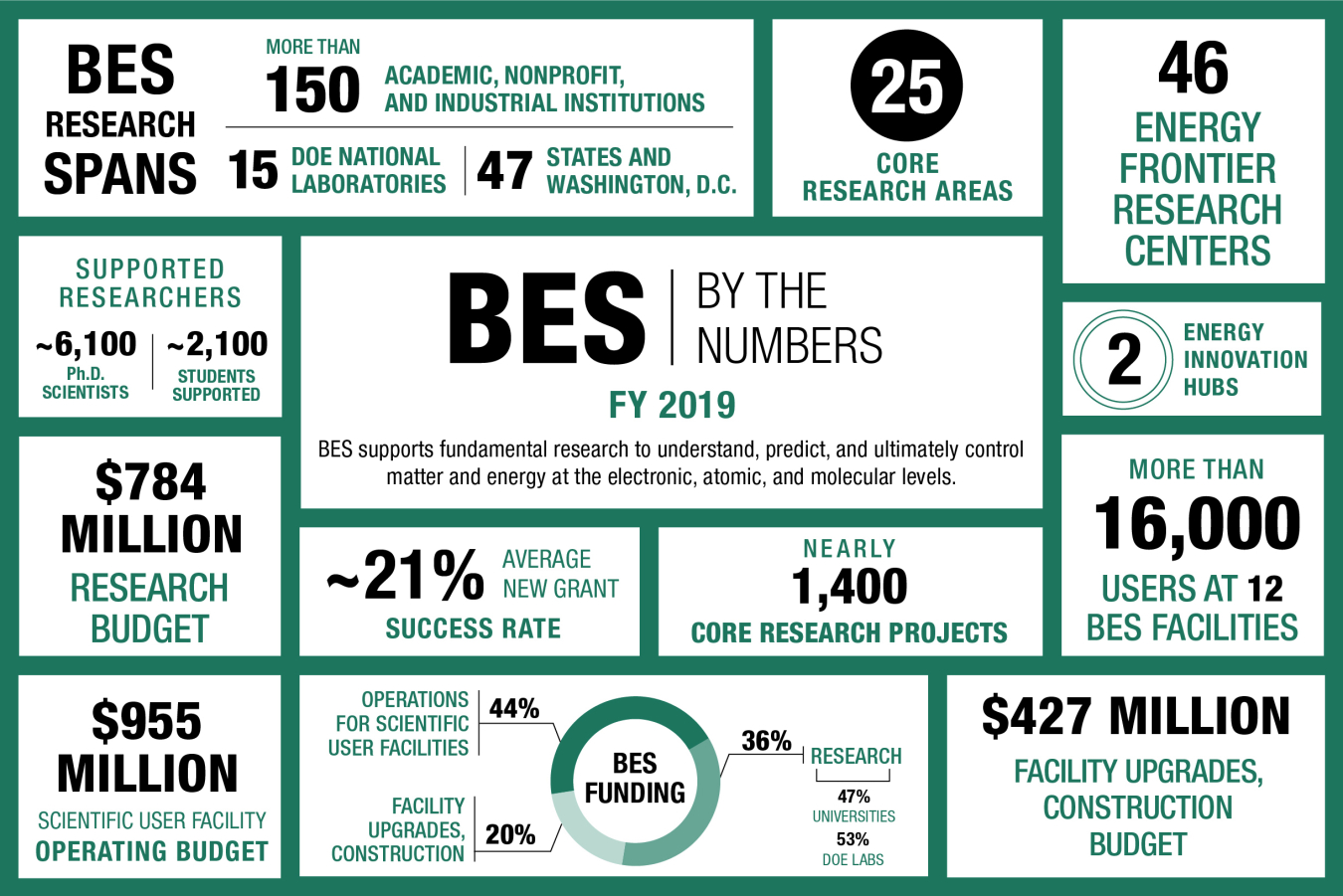
The BES program is one of the nation’s largest sponsors of research in the physical sciences. The program funds basic science at nearly 170 universities, national laboratories, and other research institutions in the U.S. BES has also built and supports a national network of major shared research facilities based at DOE national laboratories and open to all scientists. These user facilities help form the backbone of the nation’s research infrastructure. Over 16,000 scientists and engineers make use of these facilities each year.
Learn more about the Basic Energy Sciences mission and operations.

BES Subprograms
Chemical Sciences, Geosciences, and Biosciences (CSGB)
The Chemical Sciences, Geosciences, and Biosciences Division supports basic research on chemical transformations and energy flow. This research provides the groundwork for the development of new and improved processes for the generation, storage, conversion, and use of energy as well as for other applications.
Materials Sciences and Engineering (MSE)
The Materials Sciences and Engineering Division supports basic research for the discovery and design of new materials with novel properties and functions. This research creates a foundation for the development of new and improved materials for the generation, storage, conversion, and use of energy as well as for other applications.
Scientific User Facilities (SUF)
The Scientific User Facilities Division supports R&D, planning, construction, and operation of a nationwide suite of major scientific facilities. These user facilities include large x-ray light sources, neutron scattering centers, and research centers for nanoscale science. They provide state-of-the-art instrumentation to create and measure materials and chemical systems. Tens of thousands of scientists from universities, industry, and government laboratories use them each year.
Energy Frontier Research Centers (EFRCs)
The Energy Frontier Research Centers bring together teams of scientists to perform basic research with a scope and complexity beyond what is possible for individuals or small groups. These centers foster transformative scientific advances to uncover innovative solutions to difficult problems in the energy sciences..
Computational Materials and Chemical Sciences (CMS,CCS)
Computational Materials and Chemical Sciences supports teams of researchers performing basic research to develop software and databases for design of new materials and chemical processes. This research takes advantage of DOE’s current supercomputers and develops software for next-generation exascale computing systems.
Energy Innovation Hubs
The Energy Innovation Hubs mobilize large research teams to overcome major scientific barriers to development of transformative new energy technologies. The two Hubs supported by BES focus on grand challenges in energy: (1) Fuels from Sunlight and (2) Next Generation Batteries and Energy Storage.
BES Science Highlights
BES Program News
BES Research Resources
Contact Information
Basic Energy Sciences
U.S. Department of Energy
SC-22/Germantown Building
1000 Independence Avenue., SW
Washington, DC 20585
P: (301) 903 - 3081
F: (301) 903 - 6594
E: sc.bes@science.doe.gov